pynibs package¶
Subpackages¶
Submodules¶
pynibs.freesurfer module¶
This holds methods to interact with FreeSurfer ([1]), for example to translate FreeSurfer files into Paraview readable .vtk files.
References
Dale, A.M., Fischl, B., Sereno, M.I., 1999. Cortical surface-based analysis. I. Segmentation and surface reconstruction. Neuroimage 9, 179-194.
- pynibs.freesurfer.data_sub2avg(fn_subject_obj, fn_average_obj, hemisphere, fn_in_hdf5_data, data_hdf5_path, data_label, fn_out_hdf5_geo, fn_out_hdf5_data, mesh_idx=0, roi_idx=0, subject_data_in_center=True, data_substitute=-1, verbose=True, replace=True, reg_fn='sphere.reg')¶
Maps the data from the subject space to the average template. If the data is given only in an ROI, the data is mapped to the whole brain surface.
- Parameters:
fn_subject_obj (str) – Filename of subject object .hdf5 file (incl. path), e.g.
.../probands/subjectID/subjectID.hdf5fn_average_obj (str) – Filename of average template object .pkl file (incl. path), e.g.
.../probands/avg_template/avg_template.hdf5.hemisphere (str) – Define hemisphere to work on (
'lh'or'rh'for left or right hemisphere, respectively).fn_in_hdf5_data (str) – Filename of .hdf5 data input file containing the subject data.
data_hdf5_path (str) – Path in .hdf5 data file where data is stored (e.g.
'/data/tris/').data_label (str or list of str) – Label of datasets contained in hdf5 input file to map.
fn_out_hdf5_geo (str) – Filename of .hdf5 geo output file containing the geometry information.
fn_out_hdf5_data (str) – Filename of .hdf5 data output file containing the mapped data.
mesh_idx (int) – Index of mesh used in the simulations.
roi_idx (int) – Index of region of interest used in the simulations.
subject_data_in_center (bool, default: True) – Specify if the data is given in the center of the triangles or in the nodes.
data_substitute (float) – Data substitute with this number for all points outside the ROI mask
verbose (bool) – Verbose output (Default: True)
replace (bool) – Replace output files (Default: True)
reg_fn (str) – Sphere.reg fn
- Returns:
<Files> – Geometry and corresponding data files to plot with Paraview:
fn_out_hdf5_geo.hdf5: geometry file containing the geometry information of the average template
fn_out_hdf5_data.hdf5: geometry file containing the data
- Return type:
.hdf5 files
- pynibs.freesurfer.freesurfer2vtk(in_fns, out_folder, hem='lh', surf='pial', prefix=None, fs_subject='fsaverage', fs_subjects_dir=None)¶
Transform multiple FreeSurfer .mgh files into one .vtk file. This can be read with Paraview and others.
- Parameters:
out_folder (str) – Output folder.
hem (str, default: 'lh') – Which hemisphere:
'lh'or'rh'.surf (str, default: 'pial') – Which FreeSurfer surface:
'pial','inflated', …prefix (str, optional) – Prefix to add to each filename.
fs_subject (str, default: 'fsaverage') – FreeSurfer subject.
fs_subjects_dir (str, optional) – FreeSurfer subjects directory. If not provided, read from environment.
- Returns:
<File> – One .vtk file with data as overlays from all .mgh files provided
- Return type:
out_folder/{prefix}_{hem}_{surf}.vtk
- pynibs.freesurfer.make_average_subject(subjects, subject_dir, average_dir, fn_reg='sphere.reg')¶
Generates the average template from a list of subjects using the FreeSurfer average.
- Parameters:
subjects (list of str) – Paths of subjects directories, where the FreeSurfer files are located, e.g. for simnibs mri2mesh
.../fs_SUBJECT_ID.subject_dir (str) – Temporary subject directory of FreeSurfer (symlinks of subjects will be generated in there and average template will be temporarily stored before it is copied to
average_dir).average_dir (str) – path to directory where average template will be stored, e.g. probands/avg_template_15/mesh/0/fs_avg_template_15.
fn_reg (str, default: 'sphere.reg' --> ?h.sphere.reg>) – Filename suffix of FreeSurfer registration file containing registration information to template.
- Returns:
<Files> – Average template in average_dir and registered curvature files,
?h.sphere.regin subjects/surf folders.- Return type:
.tif and .reg files
- pynibs.freesurfer.make_group_average(subjects=None, subject_dir=None, average=None, hemi='lh', template='mytemplate', steps=3, n_cpu=2, average_dir=None)¶
Creates a group average from scratch, based on one subject. This prevents for example the fsaverage problems of large elements at M1, etc. This is an implemntation of [2] ‘Creating a registration template from scratch (GW)’.
References
- Parameters:
subject_dir (str) – Temporary subject directory of FreeSurfer (symlinks of subjects will be generated in there and average template will be temporarily stored before it is copied to
average_dir).average (str, default:
subjects[0]) – Which subject to base new average template on.hemi (str, default: 'lh') –
Which hemisphere:
lhorrh.Deprecated since version 0.0.1: Don’t use anymore.
template (str, default: 'mytemplate') – Basename of new template.
steps (int, default: 2) – Number of iterations.
n_cpu (int, default: 4) – How many cores for multithreading.
average_dir (str) – Path to directory where average template will be stored, e.g.
probands/avg_template_15/mesh/0/fs_avg_template_15.
- Returns:
<File> (.tif file) –
SUBJECT_DIR/TEMPLATE*.tif, TEMPLATE0.tif based on AVERAGE, rest on all subjects.<File> (.myreg file) –
SUBJECT_DIR/SUBJECT*/surf/HEMI.sphere.myreg*.<File> (.tif file) – Subject wise sphere registration based on
TEMPLATE*.tif.
- pynibs.freesurfer.read_curv_data(fname_curv, fname_inf, raw=False)¶
Read curvature data provided by FreeSurfer with optional normalization.
- Parameters:
fname_curv (str) – Filename of the FreeSurfer curvature file (e.g. ?h.curv), contains curvature data in nodes can be found in mri2mesh proband folder:
proband_ID/fs_ID/surf/?h.curv.fname_inf (str) – Filename of inflated brain surface (e.g.
?h.inflated), contains points and connectivity data of surface can be found in mri2mesh proband folder:proband_ID/fs_ID/surf/?h.inflated.raw (bool) – If raw-data is returned or if the data is normalized to -1 for neg. and +1 for pos. curvature.
- Returns:
curv – Curvature data in element centers.
- Return type:
pynibs.muap module¶
- pynibs.muap.calc_mep_wilson(firing_rate_in, t, Qvmax=900, Qmmax=300, q=8, Tmin=14, N=100, M0=42, lam=0.002, tau0=0.006)¶
Determine motor evoked potential from incoming firing rate
- Parameters:
firing_rate_in (ndarray of float [n_t]) – Input firing rate from alpha motor neurons
t (ndarray of float [n_t]) – Time axis in s
Qvmax (float, optional, default: 900) – Max of incoming firing rate [1/s]
Qmmax (float, optional, default: 300) – Max of MU firing rate [1/s]
q (float, optional, default: 8) – Min firing rate of MU [1/s]
Tmin (float, optional, default: 14) – Min MU threshold [1/s]
N (float, optional, default: 100) – Number of MU
M0 (float, optional, default: 42) – Scaling constant of MU amplitude [mV/s]
lam (float, optional, default: 0.002) – MUAP timescale of first order Hermite Rodriguez function [s]
tau0 (float, optional, default: 0.006) – Standard shift of MUAP to ensure causality [s]
- Returns:
mep – Motor evoked potential at surface electrode
- Return type:
ndarray of float [n_t]
- pynibs.muap.compute_signal(signal_matrix, sensor_matrix)¶
Determine average signal from one single muscle fibre on all point electrodes
- Parameters:
- Returns:
signal – Average signal detected all point electrodes
- Return type:
ndarray of float [n_time]
- pynibs.muap.create_electrode(l_x, l_z, n_x, n_z)¶
Creates electrode coordinates
- Parameters:
- Returns:
electrode_coords – Coordinates of point electrodes (x, y, z)
- Return type:
ndarray of float [n_ele x 3]
- pynibs.muap.create_muscle_coords(l_x, l_y, n_x, n_y, h)¶
Create x and y coordinates of muscle fibres in muscle
- Parameters:
- Returns:
muscle_coords – Coordinates of muscle fibres in x-y plane (x, y, z)
- Return type:
ndarray of float [n_muscle x 3]
- pynibs.muap.create_muscle_fibre(x0, y0, L, n_fibre)¶
Creates muscle fibre coordinates (in z-direction)
- pynibs.muap.create_sensor_matrix(electrode_coords, fibre_coords, sigma_r=1, sigma_z=1)¶
Create sensor matrix containing the inverse distances from the point electrodes to the fibre elements weighted by the anisotropy factor of the muscle tissue.
- Parameters:
electrode_coords (ndarray of float [n_ele x 3]) – Coordinates of point electrodes (x, y, z)
fibre_coords (ndarray of float [n_fibre x 3]) – Coordinates of muscle fibre in z-direction (x, y, z)
sigma_r (float, optional, default: 1) – Radial conductivity of muscle
sigma_z (float, optional, default: 1) – Axial conductivity of muscle along fibre
- Returns:
sensor_matrix – Sensor matrix containing the inverse distances weighted with the anisotropy of muscle tissue
- Return type:
ndarray of float [n_fibre x n_ele]
- pynibs.muap.create_signal_matrix(T, dt, fibre_coords, z_e, v)¶
Create signal matrix containing the travelling action potential on the fibre
- Parameters:
- Returns:
signal_matrix – Signal matrix containing the action potential values for each time step in the rows
- Return type:
ndarray of float [n_time x n_fibre]
- pynibs.muap.dipole_potential(z, loc, response)¶
Returns dipole potential at given coordinates z (interpolates given dipole potential)
- pynibs.muap.hermite_rodriguez_1st(t, tau0=0, tau=0, lam=0.002)¶
First order Hermite Rodriguez function to model surface MUAPs
- Parameters:
- Returns:
y – Surface MUAP
- Return type:
ndarray of float [n_t]
- pynibs.muap.sfap(z, sigma_i=1.01, d=5.4999999999999995e-05, alpha=0.5)¶
Single fibre propagating transmembrane current (second spatial derivative of transmembrane potential).
S. D. Nandedkar and E. V. Stalberg,“Simulation of single musclefiber action potentials” Med. Biol. Eng. Comput., vol. 21, pp. 158–165, Mar.1983.
J. Duchene and J.-Y. Hogrel,“A model of EMG generation,” IEEETrans. Biomed. Eng., vol. 47, no. 2, pp. 192–200, Feb. 2000
Hamilton-Wright, A., & Stashuk, D. W. (2005). Physiologically based simulation of clinical EMG signals. IEEE Transactions on biomedical engineering, 52(2), 171-183.
- Parameters:
t (ndarray of float [n_t]) – Time in (ms)
sigma_i (float, optional, default: 1.01) – Intracellular conductivity in (S/m)
d (float, optional, default: 55*1e-6) – Diameter of muscle fibre in (m)
v (float, optional, default: 1) – Conduction velocity in (m/s)
alpha (float, optional, default: 0.5) – Scaling factor to adjust length of AP
- Returns:
i – Transmembrane current of muscle fibre
- Return type:
ndarray of float [n_t]
- pynibs.muap.sfap_dip(z)¶
- pynibs.muap.weight_signal_matrix(signal_matrix, fn_imp, t, z)¶
Weight signal matrix with impulse response from single dipole at every location
pynibs.subject module¶
- class pynibs.subject.Subject(subject_id, subject_folder)¶
Bases:
objectSubject containing subject specific information, like mesh, roi, uncertainties, plot settings.
Notes
Initialization
sub = pynibs.subject(subject_ID, mesh)
Parameters
- idstr
Subject id.
- fn_meshstr
.msh or .hdf5 file containing the mesh information.
Subject.seg, segmentation information dictionary
- fn_lh_wmstr
Filename of left hemisphere white matter surface.
- fn_rh_wmstr
Filename of right hemisphere white matter surface.
- fn_lh_gmstr
Filename of left hemisphere grey matter surface.
- fn_rh_gmstr
Filename of right hemisphere grey matter surface.
- fn_lh_curvstr
Filename of left hemisphere curvature data on grey matter surface.
- fn_rh_curvstr
Filename of right hemisphere curvature data on grey matter surface.
Subject.mri, mri information dictionary
- fn_mri_T1str
Filename of T1 image.
- fn_mri_T2str
Filename of T2 image.
- fn_mri_DTIstr
Filename of DTI dataset.
- fn_mri_DTI_bvecstr
Filename of DTI bvec file.
- fn_mri_DTI_bvalstr
Filename of DTI bval file.
- fn_mri_conformstr
Filename of conform T1 image resulting from SimNIBS mri2mesh function.
Subject.ps, plot settings dictionary
see plot functions in para.py for more details
Subject.exp, experiment dictionary
- infostr
General information about the experiment.
- datestr
Date of experiment (e.g. 01/01/2018).
- fn_tms_navstr
Path to TMS navigator folder.
- fn_datastr
Path to data folder or files.
- fn_exp_csvstr
Filename of experimental data .csv file containing the merged experimental data information.
- fn_coilstr
Filename of .ccd or .nii file of coil used in the experiment (contains ID).
- fn_mri_niistr
Filename of MRI .nii file used during the experiment.
- condstr or list of str
Conditions in the experiment in the recorded order (e.g. [‘PA-45’, ‘PP-00’]).
- experimenterstr
Name of experimenter who conducted the experiment.
- incidentsstr
Description of special events occurred during the experiment.
Subject.mesh, mesh dictionary
- infostr
Information about the mesh (e.g. discretization, etc.).
- fn_mesh_mshstr
Filename of the .msh file containing the FEM mesh.
- fn_mesh_hdf5str
Filename of the .hdf5 file containing the FEM mesh.
- seg_idxint
Index indicating to which segmentation dictionary the mesh belongs.
Subject.roi region of interest dictionary
- typestr
Specify type of ROI (‘surface’, ‘volume’)
- infostr
Info about the region of interest, e.g. “M1 midlayer from freesurfer mask xyz”.
- regionlist of str or float
Filename for freesurfer mask or
[[X_min, X_max], [Y_min, Y_max], [Z_min, Z_max]].- deltafloat
Distance parameter between WM and GM (0 -> WM, 1 -> GM) (for surfaces only).
- add_experiment_info(exp_dict)¶
Adding information about a particular experiment.
- Parameters:
exp_dict (dict of dict or list of dict) – Dictionary containing information about the experiment.
Notes
Adds Attributes
- explist of dict
Dictionary containing information about the experiment.
- add_mesh_info(mesh_dict)¶
Adding filename information of the mesh to the subject object (multiple filenames possible).
Notes
Adds Attributes
- Subject.meshlist of dict
Dictionaries containing the mesh information.
- add_mri_info(mri_dict)¶
Adding MRI information to the subject object (multiple MRIs possible).
- Parameters:
mri_dict (dict or list of dict) – Dictionary containing the MRI information of the subject.
Notes
Adds Attributes
- Subject.mrilist of dict
Dictionary containing the MRI information of the subject.
- add_plotsettings(ps_dict)¶
Adding ROI information to the subject object (multiple ROIs possible).
Notes
Adds Attributes
- Subject.pslist of dict
Dictionary containing plot settings of the subject.
- add_roi_info(roi_dict)¶
Adding ROI (surface) information of the mesh with mesh_index to the subject object (multiple ROIs possible).
- Parameters:
roi_dict (dict of dict or list of dict) – Dictionary containing the ROI information of the mesh with mesh_index
[mesh_idx][roi_idx].
Notes
Adds Attributes
- Subject.mesh[mesh_index].roilist of dict
Dictionaries containing ROI information.
- pynibs.subject.check_file_and_format(fname)¶
Checking existence of file and transforming to list if necessary.
- pynibs.subject.create_plot_settings_dict(plotfunction_type)¶
Creates a dictionary with default plotsettings.
- Parameters:
plotfunction_type (str) –
Plot function the dictionary is generated for:
’surface_vector_plot’
’surface_vector_plot_vtu’
’volume_plot’
’volume_plot_vtu’
- Returns:
ps (dict) – Dictionary containing the plotsettings.
axes (bool) – Show orientation axes.
background_color (np.ndarray) – (1m 3) Set background color of exported image RGB (0…1).
calculator (str) – Format string with placeholder of the calculator expression the quantity to plot is modified with, e.g.: “{}^5”.
clip_coords (np.ndarray of float) – (N_clips, 3) Coordinates of clip surface origins (x,y,z).
clip_normals (np.ndarray of float) – (N_clips, 3) Surface normals of clip surfaces pointing in the direction where the volume is kept for clip_type = [‘clip’ …] (x,y,z).
clip_type (list of str) – Type of clipping:
’clip’: cut geometry but keep volume behind
’slice’: cut geometry and keep only the slice
coil_dipole_scaling (list [1 x 2]) – Specify the scaling type of the dipoles (2 entries):
coil_dipole_scaling[0]:’uniform’: uniform scaling, i.e. all dipoles have the same size
’scaled’: size scaled according to dipole magnitude
coil_dipole_scaling[1]:scalar scale parameter of dipole size
coil_dipole_color (str or list) – Color of the dipoles; either str to specify colormap (e.g. ‘jet’) or list of RGB values [1 x 3] (0…1).
coil_axes (bool, default: True) – Plot coil axes visualizing the principle direction and orientation of the coil.
colorbar_label (str) – Label of plotted data close to colorbar.
colorbar_position (list of float) – (1, 2) Position of colorbar (lower left corner) 0…1 [x_pos, y_pos].
colorbar_orientation (str) – Orientation of colorbar (
'Vertical','Horizontal').colorbar_aspectratio (int) – Aspectratio of colorbar (higher values make it thicker).
colorbar_titlefontsize (float) – Fontsize of colorbar title.
colorbar_labelfontsize (float) – Fontsize of colorbar labels (numbers).
colorbar_labelformat (str) – Format of colorbar labels (e.g.: ‘%-#6.3g’).
colorbar_numberoflabels (int) – maximum number of colorbar labels.
colorbar_labelcolor (list of float) – (1, 3) Color of colorbar labels in RGB (0…1).
colormap (str or np.ndarray) – If np.ndarray [1 x 4*N]: custom colormap providing data and corresponding RGB values
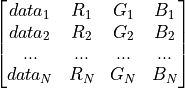
if str: colormap of plotted data chosen from included presets:
’Cool to Warm’,
’Cool to Warm (Extended)’,
’Blue to Red Rainbow’,
’X Ray’,
’Grayscale’,
’jet’,
’hsv’,
’erdc_iceFire_L’,
’Plasma (matplotlib)’,
’Viridis (matplotlib)’,
’gray_Matlab’,
’Spectral_lowBlue’,
’BuRd’
’Rainbow Blended White’
’b2rcw’
colormap_categories (bool) – Use categorized (discrete) colormap.
datarange (list) – (1, 2) Minimum and Maximum of plotted datarange [MIN, MAX] (default: automatic).
domain_IDs (int or list of int) – Domain IDs surface plot: Index of surface where the data is plotted on (Default: 0) volume plot: Specify the domains IDs to show in plot (default: all) Attention! Has to be included in the dataset under the name ‘tissue’! e.g. for SimNIBS:
1 -> white matter (WM)
2 -> grey matter (GM)
3 -> cerebrospinal fluid (CSF)
4 -> skull
5 -> skin
domain_label (str) – Label of the dataset which contains the domain IDs (default: ‘tissue_type’).
edges (BOOL) – Show edges of mesh.
fname_in (str or list of str) – Filenames of input files, 2 possibilities:
.xdmf-file: filename of .xmdf (needs the corresponding .hdf5 file(s) in the same folder)
.hdf5-file(s): filename(s) of .hdf5 file(s) containing the data and the geometry. The data can be provided in the first hdf5 file and the geometry can be provided in the second file. However, both can be also provided in a single hdf5 file.
fname_png (str) – Name of output .png file (incl. path).
fname_vtu_volume (str) – Name of .vtu volume file containing volume data (incl. path).
fname_vtu_surface (str) – Name of .vtu surface file containing surface data (incl. path) (to distinguish tissues).
fname_vtu_coil (str, optional) – Name of coil .vtu file (incl. path).
info (str) – Information about the plot the settings belong to.
interpolate (bool) – Interpolate data for visual smoothness.
NanColor (list of float) –
RGB color values for “Not a Number” values (range 0 … 1).
opacitymap (np.ndarray) – Points defining the piecewise linear opacity transfer function (transparency) (default: no transparency) connecting data values with opacity (alpha) values ranging from 0 (max. transparency) to 1 (no transparency).
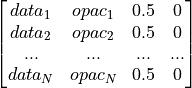
plot_function (str) – Function the plot is generated with:
’surface_vector_plot’
’surface_vector_plot_vtu’
’volume_plot’
’volume_plot_vtu’
png_resolution (float) – Resolution parameter of output image (1…5).
quantity (str) – Label of magnitude dataset to plot.
surface_color (np.ndarray) – (1, 3) Color of brain surface in RGB (0…1) for better visibility of tissue borders.
surface_smoothing (bool) – Smooth the plotted surface (True/False).
show_coil (bool, default: True) – show coil if present in dataset as block termed ‘coil’.
vcolor (np.ndarray of float) – (N_vecs, 3) Array containing the RGB values between 0…1 of the vector groups in dataset to plot.
vector_mode (dict) – Key determines the type how many vectors are shown:
’All Points’
’Every Nth Point’
’Uniform Spatial Distribution’
Value (int) is the corresponding number of vectors:
’All Points’ (not set)
’Every Nth Point’ (every Nth vector is shown in the grid)
’Uniform Spatial Distribution’ (not set)
view (list) – Camera position and angle in 3D space:
[[3 x CameraPosition], [3 x CameraFocalPoint], [3 x CameraViewUp], 1 x CameraParallelScale].viewsize (np.ndarray [1 x 2]) – Set size of exported image in pixel [width x height] will be extra scaled by parameter png_resolution.
vlabels (list of str) – Labels of vector datasets to plot (other present datasets are ignored).
vscales (list of float) – Scale parameters of vector groups to plot.
vscale_mode (list of str [N_vecs x 1]) – List containing the type of vector scaling:
’off’: all vectors are normalized
’vector’: vectors are scaled according to their magnitudes
- pynibs.subject.fill_from_dict(obj, d)¶
Set all attributes from d in obj.
- Parameters:
obj (pynibs.Mesh or pynibs.roi.ROI) – Object to fill with
d.d (dict) – Dictionary containing the attributes to set.
- Returns:
obj – Object with attributes set from
d.- Return type:
pynibs.Mesh or pynibs.ROI
- pynibs.subject.load_subject(fname, filetype=None)¶
Wrapper for pkl and hdf5 subject loader
- Parameters:
- Returns:
subject – Loaded Subject object.
- Return type:
- pynibs.subject.load_subject_hdf5(fname)¶
Loading subject information from .hdf5 file and returning subject object.
- Parameters:
fname (str) – Filename with .hdf5 extension (incl. path).
- Returns:
subject – The Subject object.
- Return type:
- pynibs.subject.load_subject_pkl(fname)¶
Loading subject object from .pkl file.
- Parameters:
fname (str) – Filename with .pkl extension.
- Returns:
subject – Loaded Subject object.
- Return type:
- pynibs.subject.save_subject(subject_id, subject_folder, fname, mri_dict=None, mesh_dict=None, roi_dict=None, exp_dict=None, ps_dict=None, **kwargs)¶
Saves subject information in .pkl or .hdf5 format (preferred)
- Parameters:
- Returns:
<File> – Subject information
- Return type:
.hdf5 file
- pynibs.subject.save_subject_hdf5(subject_id, subject_folder, fname, mri_dict=None, mesh_dict=None, roi_dict=None, exp_dict=None, ps_dict=None, overwrite=True, check_file_exist=False, verbose=False, **kwargs)¶
Saving subject information in hdf5 file.
- Parameters:
subject_id (str) – ID of subject.
subject_folder (str) – Subject folder.
fname (str) – Filename with .hdf5 extension (incl. path).
exp_dict (list of dict or dict of dict, optional) – Experiment info.
ps_dict (list of dict, optional, default:None) – Plot-settings info.
overwrite (bool) – Overwrites existing .hdf5 file.
check_file_exist (bool) – Hide warnings.
verbose (bool) – Print information about meshes and ROIs.
kwargs (str or np.ndarray) – Additional information saved in the parent folder of the .hdf5 file.
- Returns:
<File> – Subject information.
- Return type:
.hdf5 file
pynibs.tensor_scaling module¶
- pynibs.tensor_scaling.ellipse_eccentricity(a, b)¶
Calculates the eccentricity of an 2D ellipse with the semi axis a and b. An eccentricity of 0 corresponds to a sphere and an eccentricity of 1 means complete eccentric (line) with full restriction to the other axis
- pynibs.tensor_scaling.rescale_lambda_centerized(D, s, tsc=False)¶
Rescales the eigenvalues of the matrix D according to their eccentricity. The scale factor is between 0…1 a scale factor of 0.5 would not alter the eigenvalues of the matrix D. A scale factor of 0 would unify all eigenvalues to one value such that it corresponds to a isotropic sphere. A scale factor of 1 alters the eigenvalues in such a way that the resulting ellipsoid is fully eccentric and anisotropic.
- pynibs.tensor_scaling.rescale_lambda_centerized_workhorse(D, s, tsc=False)¶
Rescales the eigenvalues of the matrix D according to their eccentricity. The scale factor is between 0…1 a scale factor of 0.5 would not alter the eigenvalues of the matrix D. A scale factor of 0 would unify all eigenvalues to one value such that it corresponds to a isotropic sphere. A scale factor of 1 alters the eigenvalues in such a way that the resulting ellipsoid is fully eccentric and anisotropic
pynibs.tms_pulse module¶
- pynibs.tms_pulse.biphasic_pulse(t, R=0.0338, L=1.55e-05, C=0.0001936, alpha=1089.8, f=2900)¶
Returns normalized single biphasic pulse waveform of electric field (first derivative of coil current)
- Parameters:
t (ndarray of float [n_t]) – Time array in seconds
R (float, optional, default: 0.0338 Ohm) – Resistance of coil in (Ohm)
L (float, optional, default: 15.5*1e-6 H) – Inductance of coil in (H)
C (float, optional, default: 193.6*1e-6) – Capacitance of coil in (F)
alpha (float, optional, default: 1089.8 1/s) – Damping coefficient in (1/s)
f (float, optional, default: 2900 Hz) – Frequency in (Hz)
- Returns:
e – Normalized electric field time course (can be scaled with electric field)
- Return type:
ndarray of float [n_t]
